|
PerfExplorer 2.0 | ||||||||
| PREV PACKAGE NEXT PACKAGE | FRAMES NO FRAMES | ||||||||
See:
Description
| Interface Summary | |
|---|---|
| PerformanceAnalysisOperation | The PerformanceAnalysisOperation interface is defined as the methods all analysis operations should support. |
| PerformanceResult | This interface is defined as the methods all performance results should support. |
| Enum Summary | |
|---|---|
| DataNeeded.DataType | |
| ScalabilityResult.Measure | |
| ScalabilityResult.Scaling | |
The glue package provides the classes necessary to construct PerfExplorer scripts.
The glue package provides the classes which can be used to control PerfExplorer.
These classes all derive from two base interfaces,
PerformanceAnalysisOperation and
PerformanceResult. In addition to the
glue package, some classes from PerfDMF may also need to be included, such
as the Application,
Experiment, and
Trial classes.

The glue package provides a user-accessible programming interface,
with limited exposure to a number analysis data objects and process
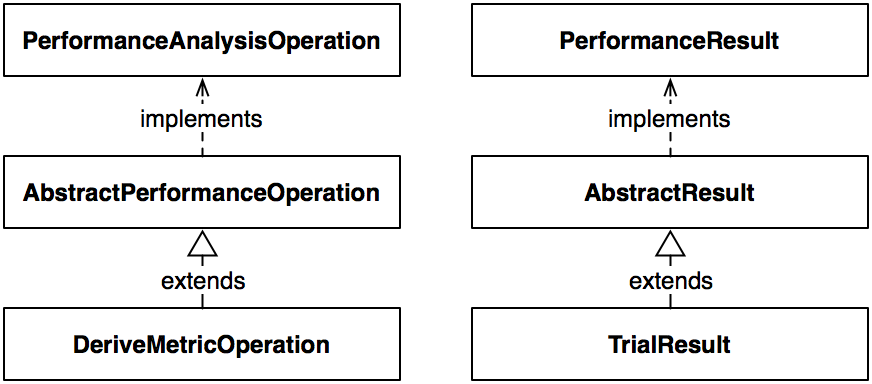
objects. As an example, take the hierarchy which constructs
the DeriveMetricOperation class.
The top level interface for the processing classes is the
PerformanceAnalysisOperation, which
defines the interface for all process objects. The interface consists of
methods to define input data, process the inputs, get output data objects,
and reset the process. The
AbstractPerformanceOperation is an
abstract implementation of the
PerformanceAnalysisOperation interface,
and provides basic internal member variables, such as the input data and
output data. Finally, the
DeriveMetricOperation class is an
example of a concrete extension of the
AbstractPerformanceOperation
class, and will take
one more more input data sets with two or more metrics each, and generate
a derived metric representing either the addition, subtraction, multiplication,
or division of one metric with the other.
Corresponding with the operation
hierarchy is the data hierarchy. At the top of the hierarchy is the
PerformanceResult
interface, which defines basic methods for
accessing the profile data within. The abstract implementation of the
interface is AbstractResult class,
which defines many internal data structures and static constants. The
TrialResult class is
an example of a class which is a concrete implementation of the
abstract class, and provides an object which holds the performance
profile data for a given trial, when loaded from PerfDMF.
Here is an example script which demonstrates the use of the package:
from edu.uoregon.tau.perfexplorer.glue import *
from edu.uoregon.tau.perfdmf import *
from java.util import *
from java.lang import *
True = 1
False = 0
def loadDB(app, exp, trial):
trial = Utilities.getTrial(app, exp, trial)
input = TrialMeanResult(trial)
return input
def loadFromDB():
# set this to your database configuration
Utilities.setSession("your_database_configuration")
# change these to the application, experiment and trials of interest
inputs.add(loadDB("application", "experiment", "trial1"))
inputs.add(loadDB("application", "experiment", "trial2"))
inputs.add(loadDB("application", "experiment", "trial3"))
inputs.add(loadDB("application", "experiment", "trial4"))
return inputs
def drawGraph(results):
# set this to the metric of interest
metric = "Time"
grapher = DrawGraph(results)
metrics = HashSet()
metrics.add(metric)
grapher.setMetrics(metrics)
grapher.setLogYAxis(False)
grapher.setShowZero(True)
grapher.setTitle("Graph of Multiple Trials: " + metric)
grapher.setSeriesType(DrawGraph.EVENTNAME)
grapher.setUnits(DrawGraph.SECONDS)
# for processor count X axis
grapher.setCategoryType(DrawGraph.PROCESSORCOUNT)
# for trial name X axis
#grapher.setCategoryType(DrawGraph.TRIALNAME)
# for metadata field on X axis
#grapher.setCategoryType(DrawGraph.METADATA)
#grapher.setMetadataField("pid")
grapher.setXAxisLabel("Processor Count")
grapher.setValueType(AbstractResult.EXCLUSIVE)
grapher.setYAxisLabel("Exclusive " + metric + " (seconds)")
grapher.processData()
def main():
print "--------------- JPython test script start ------------"
# load the data
inputs = loadFromFiles()
# extract the event of interest
events = ArrayList()
# change this to zoneLimitedGradient(PileOfScalars) as necessary
events.add("MPI_Send()")
extractor = ExtractEventOperation(inputs, events)
extracted = extractor.processData()
drawGraph(extracted)
print "---------------- JPython test script end -------------"
if __name__ == "__main__":
main()
edu.uoregon.tau.perfdmf
|
PerfExplorer 2.0 | ||||||||
| PREV PACKAGE NEXT PACKAGE | FRAMES NO FRAMES | ||||||||